Signaling with Phosphoinositides: Better than Binary
- Department of Pharmacology and Molecular Biology Program University of Colorado Health Sciences Center 4200 East Ninth Avenue, Denver, Co 80262
- Address correspondence to MO. E-mail Michael.Overduin{at}UCHSC.edu; fax 303-315-7097.
Abstract
When the outside of cell is stimulated,the inside generates a flurry of signals.Phosphates are sprinkled over lipids and proteins,where they are recognized within diverse signaling pathways.The kinases that congregate beneath the cell surface to provide the phosphate tags that mediate signaling have become major targets of new wave of drug design. Phosphoinositide signaling presents a particularly intriguing network whose many mysteries are now being unlocked.Research into protein domains that specifically recognize phosphoinositides have established the ENTH, FYVE,Phox,and pleckstrin homology domains s four cornerstones of phosphoinositide signaling.

The phosphorylation of proteins has long been regarded as the star mechanism of signal transduction. So, why focus on the
phosphorylation of phosphoinositides? Basically, because they're better than binary: The inositol ring of the phosphoinositide
head group offers multiple options for modification-and thus multiple options for recognition by the cell signaling network.
INTRODUCTION
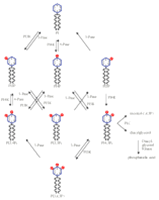
The molecules that transduce biological signals undergo modifications that are generally regarded as either “on” or “off.” Amino acid residues, for instance, are either phosphorylated or unphosphorylated. Phosphoinositides (PIs), on the other hand, can exist in any of eight permutations (Figure 1⇓). The inositol ring can be phosphorylated at the 3-, 4-, or 5- positions for a total of one, two, or three times. Some PIs reside within membrane microdomains or lipid rafts, whereas others are specifically cleaved to trigger subsequent signaling events in solution. Each PI exhibits a unique stereochemistry, and each can elicit a distinct biological response. Indeed, PI pathways constitute perhaps the most complex signaling network in the cell, and one where location and specificity are of utmost importance.
Phosphoinositide turnover. The reversible production of phosphoinositides (PIs) is catalyzed through the combined activities of a number of specific phosphatases and phosphoinositide kinases (i.e., x-Pases and PIxKs, respectively, where x designates substrate specificity with regard to the inositol ring). Cells utilize this PI network to support and distribute endocytic, exocytic, and signal transduction pathways that are, in turn, dependent upon a battery of proteins containing discrete, modular domains (viz., signaling domains) that bind to specific PIs. The figure also shows the turnover of PI4,5P2 by phospholipase C (PLC) to yield soluble inositol-1,4,5P3 and membrane-embedded diacylglycerol.
The PI signaling network offers important opportunities for therapeutic intervention, inasmuch as misread PI signals have been implicated in an array of cancers and genetic disorders. For example, major tumor suppressors and oncoproteins are responsible for the dephosphorylation and phosphorylation of PIs, respectively (reviewed in (1)). Misrecognition of PIs by mutant pleckstrin homology (PH) domains contributes to X-linked agammaglobulinemia (2), and mutations in the PI-binding site of Phox homology (PX) domains are found in chronic granulomatous disease (3–5). Mutant PI 5-phosphatases are linked with Lowe's syndrome, an X-linked disorder characterized by progressive renal failure and blindness (6). In addition, PI phosphatases have been implicated in manic depression (reviewed in (7)) and insulin sensitivity (8). The therapeutic exploitation of these findings will rest on our understanding of the critical details of PI pathways.
The function of each PI depends on when and where it is made, moved, and broken down within the cell (reviewed in (9)). Some PIs are present at fairly constant levels. The unphosphorylated form (i.e., phosphatidylinositol) comprises roughly a tenth of mammalian membrane phospholipids and is localized to cytoplasmic membrane leaflets. The most abundant monophosphorylated form is phosphatidylinositol 4-phosphate (PI4P), which is produced by PI 4-kinases resident on the Golgi apparatus so as to modulate secretion. Of the four PIs that bear a phosphate group at position 3, the most common in resting cells is the monophosphorylated form PI3P, being maintained at a sufficient concentration (~200 μM) to direct membrane traffic from endosomes and Golgi bodies to lysosomal compartments. The most abundant polyphosphorylated PI is PI4,5P2, which is enriched in regions of plasma membrane where clathrin cages are assembled for the internalization of cell surface receptors. Once the cargo is delivered within the cell, 5-phosphatases can turn over PI4,5P2 and thereby uncouple the cages from the internalized membrane. Another fate can befall PI4,5P2; it can be hydrolyzed by phospholipase C (PLC) so as to produce soluble inositol 1,4,5-trisphosphate (IP3) and membrane-bound diacylglycerol. Both products act as second messengers, boosting calcium levels and protein kinase C activity, respectively.
In contrast to the intracellular pathways that rely constitutively on PIs, levels of polyphosphorylated PIs fluctuate more dramatically at the cell surface in response to receptor activation. Such stimulation results in local bursts of PI3,4,5P3 that create plasma membrane platforms where signaling enzymes and adaptors dock; PI3,4,5P3 is then rapidly converted by 5-phosphatases into the more enduring PI3,4P2. Thus, each PI plays a dynamic role in the cell and is distributed according to the localization and activation of the kinases, phosphatases, and lipases responsible for its production and turnover.
The enzymes that underlie PI signaling and distribution, in turn, possess specific domains (i.e., “signaling domains”) through which, along with lipid anchors, they are localized—and thereby regulated—with respect to intracellular membranes. For example, the kinase that produces PI3P is recruited to endocytic membranes, where it becomes activated, by a myristoylated serine/threonine kinase and an endosomal GTPase. Subsequently, proteins that possess domains that bind to PI3P (e.g., FYVE (10–12) or PX domains (3–5, 13, 14)) are recruited to the membrane to participate in endocytosis and signaling, in some cases by acting as enzymes that metabolize PI3P. Similarly, at the cell surface, the activation of receptors results in the phosphorylation of lipids and tyrosine residues that attract signaling enzymes, such as kinases and phosphatases that contain C2 and src homology 2 (SH2) domains. Some of these enzymes have broad PI specificities that differ significantly in vitro and in vivo. Thus, their PI-recognizing domains are believed to contribute significantly to their biological specificity and to the cellular organization of the PI network (Figure 2⇓).
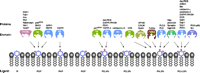
Phosphoinositide recognition domains. Signaling domains that have been demonstrated to bind phosphoinositides are shown as colored shapes with a schematized binding site. Arrows indicate the preferred ligand of the given signaling domain, and proteins that contain the given signaling domain are listed above each corresponding colored shape.
THE ENTH DOMAIN
Several proteins that assemble clathrin lattices (e.g., AP180, CALM, and Epsin) contain a PI-binding module, named the Epsin N-Terminal Homology (ENTH) domain (15), that precedes specific endocytic motifs recognized by clathrin and its associated proteins. These ENTH domains undergird the invaginating plasma membrane through PI4,5P2 interactions, and the clathrin coat connects to this scaffold (16, 17). Thus, ENTH domains provide an important link to the membrane for the system of proteins that act as ushers of clathrin-mediated endocytosis.
Based on structural studies of ENTH domains, two distinct modes of binding have been hypothesized to account for PI4,5P2 recognition. Itoh and colleagues suggest that an unstructured N terminus folds back onto a pair of helices to form the PI4,5P2 binding site of Epsin (17). How this unconventional binding site could provide ligand specificity, however, is unclear. Moreover, the solution structure (18) that provides the basis for this hypothesis differs significantly from the crystal structure of a longer sequence of the same ENTH domain (19). Nonetheless, this putative PI4,5P2 site does include three basic residues that have been implicated through mutagenesis studies as crucial to interaction with PI4,5P2 (17).
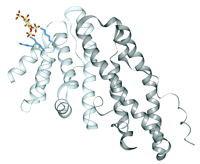
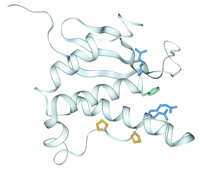
A different PI4,5P2 binding mechanism is suggested by crystal structures of the ENTH domains of AP180 and CALM (16, 20). The CALM structure reveals that the PI4,5P2 head group is perched on a positively charged apex, as shown in Figure 3⇓ (16). The 4- and 5-phosphates contact three lysines and a histidine, leaving the rest of the ligand exposed. This unusually exposed binding site could not provide stereospecificity, and may be consistent with the domain's ability to weakly bind other PIs. Alternatively, membrane docking could alter the PI binding mode, or the N terminus, which is disordered in both crystal structures (16, 20), could become engaged as suggested by Itoh and colleagues (see above; (17)).
Structure of the ENTH domain of CALM bound to the PI4,5P2 head group. A gray backbone ribbon depicts the nine α-helices of the CALM structure; the lighter shade indicates the core ENTH domain (16). The side chains of the three lysines (blue) and the histidine (green) that coordinate the phosphorylated inositol (yellow and red) are also shown. (See text for details.)
It is peculiar that the ENTH domains of AP180 and CALM are among the largest PI-binding domains and yet appear to contain the smallest PI binding sites. Thus, in addition to PI4,5P2 binding, other functions may exist for ENTH domains. For example, CALM's interactions with clathrin appear to involve this domain (21), and Epsin also interacts with transcription factors through its ENTH domain (19). The structurally similar domains of β-catenin and karyopherin interact with a variety of other proteins. The VHS domain, another relative of the ENTH domain, contains a basic surface that may bind membrane lipids (22, 23) or acidic protein sequences of receptors that sort enzymes to the lysosome (24, 25). Thus, ENTH domains appear to engage proteins as well as phospholipids, although the relationship between the two types of ligands remains to be explored.
THE FYVE DOMAIN
The specific recognition of PI3P by FYVE domains was discovered in 1998 by three research groups (10, 12). This 65-residue module is known to exist in thirty-seven human proteins (26) and is named after four proteins (i.e., Fab1, YGLO23, Vps27, and EEA1) (27). PI3P acts to tether some of these proteins, by means of their FYVE domains, to endocytic membranes (Figure 2⇑), as exemplified by the mislocalization of early endosomal antigen 1 (EEA1) when key residues in its FYVE domain are mutated (10, 28). This membrane fusion protein is also an autoantigen associated with subacute cutaneous systemic lupus erythematosus (29) and several neurological diseases (30). Its FYVE domain binds PI3P-containing membranes with high affinity (Kd = 50 nM) (31) and is the first PI-binding domain shown to insert itself into the membrane bilayer (32). The anchoring of EEA1 at the endosome involves insertion of a hydrophobic loop into the membrane interior, specific binding of the PI3P head group (32), and parallel EEA1 dimerization, perhaps to enhance PI3P avidity (28, 33). In addition, the interaction of EEA1's FYVE domain with endosomes may be modulated by the GTPase Rab5 (34). Thus, EEA1 is held to the membrane by multiple interactions that together provide the specificity, affinity, and regulation needed for proper membrane fusion by soluble N-ethylmaleimide-sensitive factor attachment protein receptors (SNAREs).
Several other FYVE domain–containing proteins are involved in vesicular traffic (reviewed in (35)). For example, the fusion of endosomes relies on the FYVE domain of Rabenosyn-5 to bind PI3P, thereby modulating SNARE complex formation with Rab5-GTPase and Sec1 proteins. Similarly, the Hrs protein mediates the maturation of endosomes, also by interactions with SNARE proteins, and although its FYVE domain binds to PI3P-containing membranes tightly, multiple FYVE domains may cooperate for stable membrane attachment (22). Transcription factors Smad2 and Smad3 are recruited to the activated transforming growth factor β receptor by an adaptor protein, known as SARA (for Smad anchor for receptor activation), that is localized to endosomes presumably by its FYVE domain. The Fgd protein is implicated in faciogenital dysplasia, an X-linked developmental disease, and belongs to a family of proteins that regulate the actin cytoskeleton by acting as guanine nucleotide exchange factors. The FYVE and PH domains of Fgd proteins are thought to help form lamellipodia and filipodia (i.e., fine actin-based extensions involved in cell movement). Enzymes that possess FYVE domains include a PI3P 5-kinase known as PIKfyve (Fab1 in yeast) that regulates traffic to the lysosomal compartment. FYVE domains are also found in several relatives of myotubularin, a PI 3-phosphatase implicated in the X-linked muscle disease myotubular myopathy.
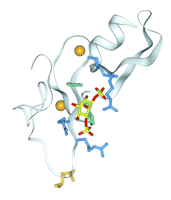
The structure and specificity of the FYVE domain are more conserved than those of other PI-binding domains. The FYVE domains of EEA1 (32), Hrs (22), and Vps27 (36) consist of a pair of small β-sheets and a short α-helix. Stabilization is provided by two zinc ions, each coordinated by a cluster of four cysteines (Figure 4⇓). The PI3P head group fits into a shallow groove next to the first β-strand of the EEA1 FYVE domain (32) in a manner similar to that predicted for Vps27 (36). A hallmark of its sequence, the basic RRHHCR motif, is central to coordinating both phosphates and zinc. The 1- and 3-phosphates are positioned near pairs of conserved basic residues, and the two histidines are packed against the inositol ring (Figure 4⇓). An exposed hydrophobic loop near the 1-phosphate inserts into membrane-like micelles (28, 32, 36). An alternative binding mode has been proposed for the tandem FYVE and VHS domains of Hrs, which crystallize as a dimer that partly occludes the canonical PI3P-binding site (22).
Structure of the PI3P-bound FYVE domain of EEA1. The ribbon indicates the four β-strands and the C-terminal helix; the two zinc ions are shown as orange spheres. The side chains of an exposed threonine and valine that insert into the membrane are shown in orange at the bottom. The pairs of arginines (blue) that recognize the 3- and 1-phosphates of the inositol ring (yellow and red) are drawn above and below the two conserved histidines (green), respectively. (See (32).)
A structural mechanism has been proposed for endosome docking that involves the EEA1 protein (32). Its FYVE domain dips a membrane insertion loop nonspecifically and weakly into the membrane bilayer. This insertion may render the PI3P-binding site slightly more accessible as the domain skates across the membrane surface. Upon binding the PI3P head group, the FYVE domain assumes a more globular conformation that sits more stably in the endosomal membrane. The association of EEA1 onto endosomal membranes is regulated by Rab5, the GTPase that facilitates SNARE-mediated membrane fusion. Although the details of the ancillary protein interactions that involve EEA1 may not be generalizable, other members of the FYVE family are likely to share the bilayer penetration and PI3P recognition mechanisms.
THE PHOX HOMOLOGY DOMAIN
The Phox homology (PX) domain is a newly recognized element in the armory of PI recognition modules. This 120-residue domain was identified in 1996 and is named after the phagocyte oxidase that provides a defense against microbes (37). Over fifty human proteins, many of them membrane-localized, contain PX domains (26). In order to identify a ligand, a battle was recently waged with candidate PIs being assayed by cells, columns, filters, NMR tubes, and multi-well plates. A slew of papers has revealed that PI3P is the most common ligand of PX domains (3–5, 13, 14). The recognition of PIs by PX domains is now established for two NADPH oxidase subunits, a PI 3-kinase, several sorting nexins, and a SNARE (Figure 2⇑). Other prominent PX domain–containing proteins include phospholipases and protein kinases, whose unique roles in the cell will bring new insights to PX domain–mediated signaling.
The first PX domain–containing protein to be ascribed the specific recognition of PI3P was Vam7 (3). This SNARE functions within the yeast vacuolar membrane to usher in deliveries of vesicular cargo (38); the Vam7 PX domain is essential for targeting this SNARE to the vacuole (3). Localization of Vam7 to the vacuolar membrane is rapidly lost when PI3P production is halted in cells expressing temperature-sensitive PI 3-kinase. Moreover, the amount of Vam7 recruited to the vacuole membrane in vivo is tightly coupled to the local concentration of PI3P, and depends on the integrity of endocytic trafficking pathways. Thus, the PX domain localizes Vam7 near its PI3P-enriched site of action (i.e., the vacuole) so that its α-helical region can partner with other SNAREs on the vacuole to engage and receive vesicles.
Based on the Vam7 studies, PI3P binding appears to be mediated by a pair of basic motifs that are highly conserved in PX domains (3). The RRφ motif, consisting of Arg-Arg-Tyr/Phe residues, is functionally important, because substitution of the relevant tyrosine residue in Vam7 abolishes both vacuolar localization in vivo and PI3P binding in vitro. Another motif that includes a polar, arginine, and basic residue (pR+) is conserved near a loop that interacts with lipid micelles. Thus, the RRφ and pR+ motifs and the micelle interaction element may act together to tether the Vam7 PX domain to PI3P-containing membranes.
The sorting nexins are a large family of proteins implicated in the trafficking of activated growth factor receptors. The recruitment of sorting nexin 3 to endosomes depends on PI 3-kinase activity, and several sorting nexins bind specifically to PI3P (13). Sorting nexin 3 is targeted to endosomal PI3P pools by its PX domain and requires the integrity of its conserved RRφ motif. However, the PX domain of Vps5, a yeast sorting nexin, binds only weakly and nonspecifically to PIs in vitro, and the PX domain of the Cpk PI 3-kinase specifically interacts with PI4,5P2 (14). Thus, although PI3P appears to be the most common PX ligand, the PI specificities and affinities of some PX domains have diverged considerably.
Phagocytic cells employ the NADPH oxidase system to destroy microorganisms by generating reactive oxygen species. Individuals with defects in NAPDH oxidase subunits are compromised in their defense against pathogens, and suffer from chronic granulomatous disease. Some of the oxidase subunits are cytosolic until an activation event moves them to the plasma membrane where the catalytic subunits reside. PI kinases and phosphatases are required for membrane localization of the complex formed by the p40phox and p67phox subunits; specifically, they generate the PI3P recognized by the PX domain of the p40phox subunit (4, 5). In contrast, the PX domain of p47phox binds preferentially to PI3,4P2, but also interacts with PI3P, PI3,5P2 and PI3,4,5P3 in vitro (5). Because PI3,4P2 is a relatively rare lipid in vivo, the physiological ligand of the p47phox PX domain remains uncertain. Nonetheless, it now appears that p40phox and p47phox subunits are targeted by their PX domains to PI-enriched membranes. A point mutation in the RRφ motif of the p47phox PX domain is linked to chronic granulomatous disease (3–5), suggesting that microbial defenses are critically undermined by subunit mislocalization that renders the complete NADPH oxidase unassembled and impotent.
A structure of the ligand-free p47phox PX domain has recently been solved, revealing an antiparallel β-sheet followed by an α-helical lobe (Figure 5⇓ and (39)). The Vam7 PX structure consists of a similar topology and appears to undergo conformational changes upon PI3P binding (3). The predicted binding elements define a deep cleft, with the RRφ and pR+ motifs positioned together between two helices to coordinate the PI head group. The pR+ motif is nearest the membrane interaction element, and is thus best positioned to coordinate the 1-phosphate. Moreover, a binding site for SH3 domains is partly exposed in the helical lobe of the p47phox PX domain (39). This proline-rich motif is adjacent to the membrane interaction element identified in Vam7. Consequently, the binding of a PX domain to PI-containing membranes could dislodge the polyproline helix and thereby provide better access to SH3 domains. Such instances of simultaneous binding of a PI and an SH3 domain could further restrict the membrane localization of PX proteins. Alternatively, the interaction of PX and SH3 domains may lock the protein into an autoinhibited state, with PI-containing membranes providing the key for activation. With the PX domain distracted by a PI, the SH3 domain of an NADPH oxidase subunit would then be free to interact with other NADPH oxidase subunits and assist in superoxide production.
Structure of the PX domain of p47phox. The backbone ribbon depicts the three-stranded antiparallel β-sheet followed by a bundle of three α-helices and short 3/10 helix (39). A pair of proline side chains is shown in yellow to indicate the polyproline type II helix recognized by the SH3 domain. The predicted PI binding elements (3) that form an accessible basic pocket include the arginine (blue) and phenylalanine (green) side chains of the RRφ motif that precedes the first α-helix and pR+ motif in the second α-helix. (See text for details.)
THE PLECKSTRIN HOMOLOGY DOMAIN
The pleckstrin homology (PH) domain is the most thoroughly characterized of the PI recognition modules. Since its discovery in 1993 (40, 41), its 120-residue sequence has been found in several hundred human proteins (26). However, evolution has toyed extensively with these modules, pushing their sequence similarities to the barely detectable level, even inserting sets of Src homology domains into their loops. PH domains have been adopted by a wide range of protein architectures, and they fulfill a variety of roles involved in signaling, cytoskeletal organization, membrane trafficking, and lipid modification. As a consequence of this versatility, numerous classification schemes and functional predictions have been confounded.
The current consensus is that a significant minority of PH domains bind specifically and tightly to one PI, and that the majority associate weakly and promiscuously with several phospholipids. Specific recognition of six PIs has been demonstrated to date (Figure 2⇑). The most common PH target appears to be PI3,4,5P3. This lipid is bound tightly by the PH domains of proteins recruited to the plasma membrane, including Btk (i.e., a member of the Tec family of tyrosine kinases), Grp1 (i.e., a GTP-GDP exchange factor), and several GTPase-activating proteins (Figure 2⇑). On the other hand, other protein kinases and the DAPP1 adaptor bind well to both PI3,4,5P3 and PI3,4P2 through their PH domains. Such dual specificity serves to park proteins on the membrane for longer periods of time after the initial spike of PI3,4,5P3, generated upon receptor engagement, has been converted into the longer-lasting PI3,4P2. The functions of the PH family of proteins have been thoroughly reviewed elsewhere (9, 42).
A handful of PH domains appear to bind monophos-phorylated PIs such as PI3P and PI4P specifically (Figure 2⇑), whereas PI3,5P2 is targeted by a novel centaurin that resembles GTP-activating proteins (43). These diverse in vitro PI specificities considerably extend the PI signaling network. However, the biological interactions of these proteins remain to be explored, and may be complicated by competition. The local concentration of each PI in any given membrane compartment can fluctuate greatly, sometimes with dramatic consequences. For example, after PLCδ localizes to the plasma membrane through the interaction of its PH domain with PI4,5P2, its catalytic domain processively cleaves IP3 head groups from surrounding PI4,5P2 molecules in the membrane. The resulting increase in soluble IP3, which binds to the enzyme's PH domain much more tightly than does membrane-embedded PI4,5P2, thereby dislodges PLCδ from the membrane. The promiscuous dynamins present another problem. These GTPases assemble on PI4,5P2-containing membranes, helping to pinch budding vesicles off the plasma membrane during clathrin-mediated endocytosis. However, the isolated dynamin PH domain binds PI4,5P2 only weakly in vitro. The solution in vivo thus appears to entail teamwork, with dynamin proteins oligomerizing around the neck of the budding vesicle so that their collective PH domains attain the avidity needed to attach to the PI4,5P2 pool in the membrane.
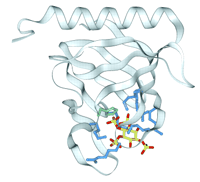
Despite its sequence diversity and functional plasticity, the PH domain's structure is highly conserved. The fold consists of a seven-stranded β-sandwich capped by an amphipathic C-terminal α-helix (Figure 6⇓). A set of variable loops is offered by a splayed corner of the β-sandwich and presents a positively charged pocket that binds PIs (44). Subsequent structures have revealed how PI head groups are bound within the PH domains of Btk, DAPP1, Grp1, PLCδ, and spectrin (2, 45–48). In each case, the inositol ring is bound in the same position, with its phosphates pinned down by hydrogen bonds to lysine and arginine side chains in the variable loops. The phosphates distal to the 1-position are the most intimately buried in the pocket, and the 1-phosphate is positioned further from the protein to help insert the acyl chains into the membrane (48). Specificity is determined in different ways. The Btk PH domain recognizes PI3,4,5P3 by an extended loop that joins β-strands 1 and 2 and wraps around the 5-phosphate (2). On the other hand, the PI3,4,5P3 specificity of the Grp1 PH domain is provided by a unique β-hairpin that extends from strands 6 and 7 (Figure 6⇓) (45, 47). The dual specificity of the DAPP1 PH domain for PI3,4P2 and PI3,4,5P3 is attributed to a lack of side chain hydrogen bonds to the 5-phosphate that would otherwise provide discrimination (45). Nonspecific PH domains such as those of spectrin exhibit fewer conserved hydrogen bonds to phosphates (46). Although the current PH structures have illuminated the binding of the PI head group, the murkier membrane interactions are also important, with a swath of positive charge surrounding each PI pocket, which is believed to interface with the membrane surface.
Structure of the PI3,4,5P3-bound PH domain of Grp1. The conserved seven stranded β-barrel fold of the PH domain is augmented by an additional hairpin shown at the lower left corner of the backbone ribbon. The inositol ring contains phospates at the 1-, 3-, 4-, and 5- positions and is bound by arginine and lysine side chains (blue) and a histidine (green), with the novel hairpin enclosing the 5-phosphate. (See (45).)
Embellishments of the PH domain fold have been discovered that mediate protein interactions (reviewed in (42)). The Dbl homology domain is an α-helical bundle that precedes some PH domains and binds nucleotide-free forms of Rho GTPases, possibly in an IP3-dependent fashion. The Tec homology domain is a zinc-binding appendage near the tail of the PH domain of Btk kinases, which helps to bind G proteins. Interactions with G proteins also appear to be mediated by a C-terminal helical extension of the β-ARK PH domain. In addition, the PH fold has been adopted by several other protein interaction domains including Ena/VASP homology, PTB, and Ran binding domains, some of which may also bind PIs. Thus, individual PH domains, like other PI-binding modules, localize proteins to specific membrane compartments through a combination of interactions with PIs as well as with other proteins.
OTHER PHOSPHOINOSITIDE-BINDING DOMAINS
For those domains that we have discussed above, PI recognition is the general rule. For other domains, specific PI interactions can be the exception. In particular, C2, SH2, and tubby domains bind polyphosphorylated PIs with little stereospecificity. Nonetheless, these interactions are important, attracting diverse cytosolic proteins to the plasma membrane in response to receptor activation.
The C2 domain recognizes acidic phospholipids and was identified as the second conserved region of protein kinase C (49). Around 200 human proteins are known to contain these 130-residue domains, including phospholipases, PI 3-kinases, protein kinases, Ras GTPase activating proteins, and synaptotagmins (26). The structures of several C2 domains have been solved, revealing an eight-stranded β-sandwich with flexible loops forming a lip of the barrel that coordinates calcium (reviewed in (50)). C2 domains bind phospholipid membranes in either calcium dependent or independent modes, and some also interact with intracellular proteins. However, there are several proteins, most notably rabphilin3A and synaptotagmin II, whose C2 domains target PIs. Synaptotagmin II binds to polyphosphorylated inositols with submicromolar affinity through one of its C2 domains, and prefers PI4,5P2 and PI3,4,5P3 (51). Its clusters of basic residues may mediate binding and are loosely conserved in rabphilin3A, a protein that binds the same PIs (52). Synaptotagmin interacts with its PI ligands to contribute to the exocytic release of neurotransmitters into the synapse, whereas rabphilin3A's C2 domains may, through Rab3 GTPases, regulate the secretion of synaptic vesicles.
The SH2 domains of the Src oncoprotein and PI3K p85 subunit have been shown to interact with PI3,4,5P3. Moreover, this noncanonical interaction competes with the more typical ligands of SH2 domains, namely, phosphotyrosine-containing protein sequences (53). Furthermore, a regulatory role has been suggested for a similar interaction of the SH2 domain of PLCγ with PI3,4,5P3, which enhances the enzyme's activity in vitro (54, 55). However, the structural basis of these SH2-mediated interactions with PIs remains unclear.
The newest recruit to the PI-binding family is the tubby domain. This module is thought to bind PI4,5P2 in vivo and has been found at the C terminus of four human proteins (56). Its sequence of 260 residues forms a unique β-barrel that holds a helix inside. The outside of the tubby barrel contains a conserved, basic pocket that can accommodate not only PI4,5P2, but also PI3,4P2 and PI3,4,5P3. Santagata and coworkers propose that PI4,5P2, which is the most common of these PIs in the plasma membrane, constrains Tubby to the membrane surface until phospholipases cleave the ligand, allowing the protein to float away to the nucleus to regulate transcription of genes that influence obesity and energy balance.
CONCLUSIONS
The discovery that PIs provide membranes with the properties needed to attract a diversity of protein domains has fueled an explosion in the literature of signal transduction and membrane traffic. What is emerging is an extensive PI network that relies on subtle differences in inositol stereochemistry to direct the flow of membrane compartments and associated proteins through the cellular maze. It is now clear that ENTH, FYVE, PH, and PX domains direct hundreds of cytoplasmic proteins to PI-enriched membranes. The interactions between PIs and these domains can serve to localize proteins transiently, or can participate in more stable membrane docking in cases where they are reinforced by hydrophobic insertions into the bilayer, by protein multimerization, or by other proteins associated with the membrane. This PI network is superimposed on the traditional signaling pathways that rely on intracellular protein modifications and interactions. It is through the interface of the PI and protein modification networks that proteins find their rightful homes on membranes. Thus, the principles of signal transduction and cellular localization are closely intertwined, and understanding their interplay will be essential for homing in on the mechanisms of disease and therapeutics.
Acknowledgments
The authors would like to thank Trey K. Sato, Christopher G. Burd, and Scott D. Emr for discussions, and Daisuke Kohda for the p47phox PX domain coordinates. We apologize to the many authors whose work could not be directly cited due to page restrictions.
- © American Society for Pharmacology and Experimental Theraputics 2001
References

Michael Overduin, Ph.D., is an Assistant Professor of Pharmacology and Biochemistry and Molecular Genetics, Pew Scholar, and faculty member of the Programs in Biomolecular Structure, Molecular Biology, and University of Colorado Cancer Center. Matthew L. Cheever (right) is a senior graduate student in the Molecular Biology Program. Tatiana G. Kutateladze, Ph.D., is an Instructor in the Department of Pharmacology and faculty member of the Program in Biomolecular Structure.